【3D Genome】 應用電腦視覺於TAD拓撲結構域之預測創意發想

由於Hi-C 視覺化之後,可視為一個二維矩陣,類似於灰階影像。而拓撲結構域Topologically associating domains (TAD),被定義為相較其外部,它的內部有高度交互作用的一個方形區域。因此可以利用先將Hi-C矩陣轉換到影像空間之後,再利用常見的電腦視覺的技巧把TAD框出來,類似傳統找ROI的處理技巧來做到這件事。
方法設計
下圖是設計的流程,大致上按照傳統電腦視覺的邏輯去搭建
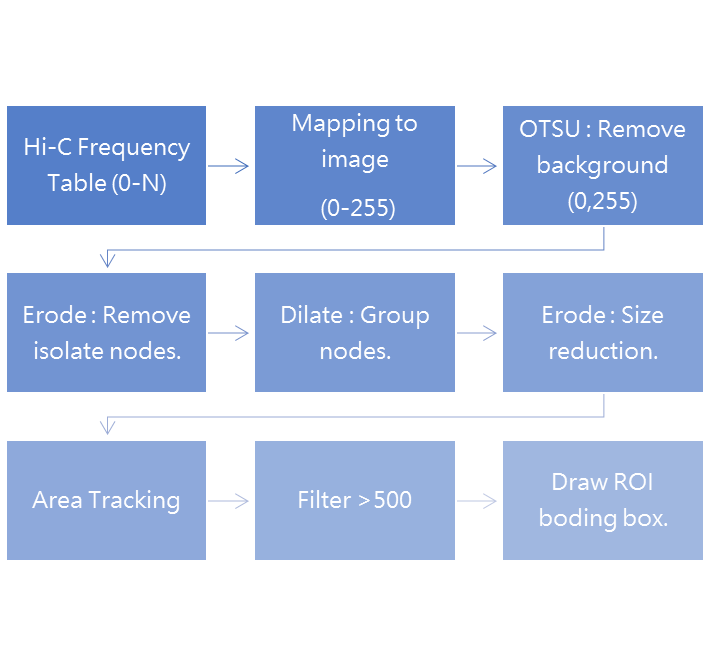
實驗討論
實作的效果如下圖

,最終call出 的TAD由綠色框線所標示
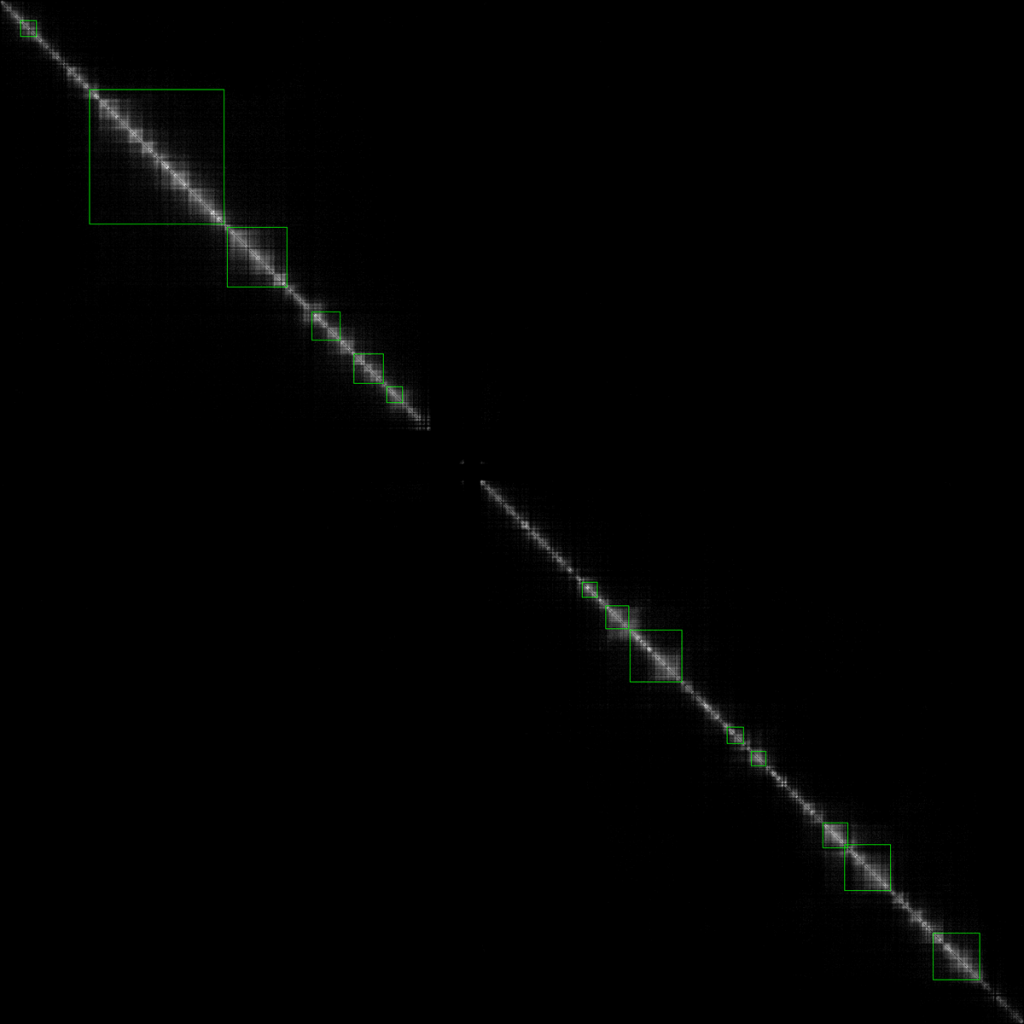

經過一些簡單的實驗之後知道,把HiC矩陣轉換成視覺圖的參數需要自動化,並且引入一個類似感知機的方程式,類似下圖
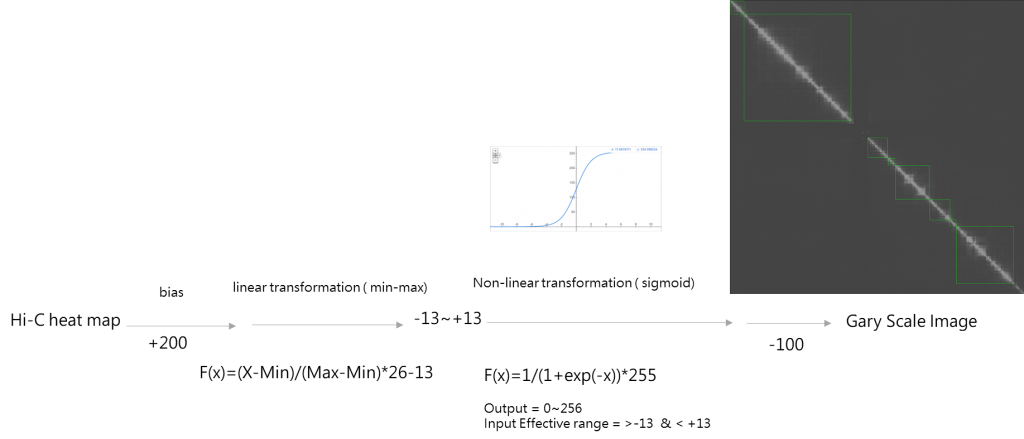
實作程式碼
HiCCVTAD.java
package com.whuang022.bio;
import org.bytedeco.javacpp.opencv_core.Mat;
import org.bytedeco.javacpp.opencv_core.MatVector;
/**
* HiC TAD DTO Class
* @author whuang022
*/
public class HiCCVTAD{
public Mat TADImage;
public MatVector TADRects;
}HiCCV.java
package com.whuang022.bio;
import org.bytedeco.javacpp.BytePointer;
import org.bytedeco.javacpp.opencv_core;
import static org.bytedeco.javacpp.opencv_core.IPL_DEPTH_8U;
import org.bytedeco.javacpp.opencv_core.IplImage;
import org.bytedeco.javacpp.opencv_core.Mat;
import org.bytedeco.javacpp.opencv_core.MatVector;
import org.bytedeco.javacpp.opencv_core.Scalar;
import static org.bytedeco.javacpp.opencv_core.cvCreateImage;
import static org.bytedeco.javacpp.opencv_core.cvGetSize;
import static org.bytedeco.javacpp.opencv_imgproc.COLOR_GRAY2BGR;
import static org.bytedeco.javacpp.opencv_imgproc.CV_CHAIN_APPROX_NONE;
import static org.bytedeco.javacpp.opencv_imgproc.CV_RETR_EXTERNAL;
import static org.bytedeco.javacpp.opencv_imgproc.CV_THRESH_BINARY;
import static org.bytedeco.javacpp.opencv_imgproc.CV_THRESH_OTSU;
import static org.bytedeco.javacpp.opencv_imgproc.boundingRect;
import static org.bytedeco.javacpp.opencv_imgproc.cvDilate;
import static org.bytedeco.javacpp.opencv_imgproc.cvErode;
import static org.bytedeco.javacpp.opencv_imgproc.cvtColor;
import static org.bytedeco.javacpp.opencv_imgproc.findContours;
import static org.bytedeco.javacpp.opencv_imgproc.rectangle;
import static org.bytedeco.javacpp.opencv_imgproc.threshold;
/**
* HiC JavaCV Process Class
* @author whuang022
*/
public class HiCCV {
public static Mat HiCtoMat (HiC hic,int shiftVal){
IplImage image = IplImage.create(hic.getRowSize(),hic.getColSize(), IPL_DEPTH_8U, 1);
Mat mImage=new Mat(image);
for(int i=1;i<hic.getRowSize();i++){
for(int j=1;j<hic.getColSize();j++){
BytePointer bytePointer = mImage.ptr(i,j,0);
double val=hic.get(i,j);
double color= ((val - hic.getMin()) * (1/( hic.getMax() - hic.getMin()) * 255)); //linear mapping HiC to Gay Scale Image (0-255)
bytePointer.putInt((int)color+shiftVal);
}
}
return mImage;
}
public static HiCCVTAD HiCTADCallingMorphology (HiC hic,int shiftVal,int min){
Mat HiCMat;
Mat otsu = new Mat();
IplImage otsuIp ;
IplImage colseIp ;
Mat HicMatOut = new Mat();
HiCMat=HiCCV.HiCtoMat(hic,shiftVal);
MatVector contours = new MatVector();
MatVector outcontours = new MatVector();
threshold(HiCMat, otsu, 10, 255, CV_THRESH_OTSU + CV_THRESH_BINARY); // OTSU to Mark Acctivate Zone
otsuIp = new IplImage(otsu);
colseIp= cvCreateImage(cvGetSize(otsuIp), IPL_DEPTH_8U, 1);
//Opening Operation
cvErode(otsuIp, colseIp, null, 1);
cvDilate(colseIp, colseIp, null, 3);
cvDilate(colseIp, colseIp, null, 1);
Mat colse=new Mat(colseIp);
findContours(colse, contours,CV_RETR_EXTERNAL, CV_CHAIN_APPROX_NONE);
cvtColor(HiCMat,HicMatOut, COLOR_GRAY2BGR);
for(int i=0;i<contours.size();i++){
opencv_core.Rect mr = boundingRect(contours.get(i));
if(mr.area()>min){
rectangle(HicMatOut, mr, new Scalar(0, 255, 0, 1));
outcontours.put(mr);
}
}
HiCCVTAD out =new HiCCVTAD();
out.TADImage=HicMatOut;
out.TADRects=outcontours;
return out;
}
}
HiC.java
package com.whuang022.bio;
import java.io.BufferedReader;
import java.io.FileNotFoundException;
import java.io.FileReader;
import java.io.IOException;
import java.util.ArrayList;
/**
* HiC Data Reader Class
* @author whuang022
*/
public class HiC {
private ArrayList<ArrayList<Double>> data = new ArrayList<ArrayList<Double>>();
private double maxVal;
private double minVal;
public int getRowSize(){
return data.size();
}
public int getColSize(){
return data.get(0).size();
}
public double get(int row,int col){
return data.get(row).get(col);
}
public double getMax(){
return maxVal;
}
public double getMin(){
return minVal;
}
public HiC(String filePath) throws FileNotFoundException, IOException{
BufferedReader reader = new BufferedReader(new FileReader(filePath));
maxVal=Double.MIN_VALUE;
minVal=Double.MAX_VALUE;
String line ;
while((line=reader.readLine())!=null){
String[] rowData = line.split(" ");
ArrayList<Double> row=new ArrayList<>();
for (String rowData1 : rowData) {
double val=Double.parseDouble(rowData1);
row.add(val);
if(val>maxVal){
maxVal=val;
}
if(val<minVal){
minVal=val;
}
}
data.add(row);
}
}
}
實驗結果
最終的結果如下,已經可以抓出大概的TAD區域
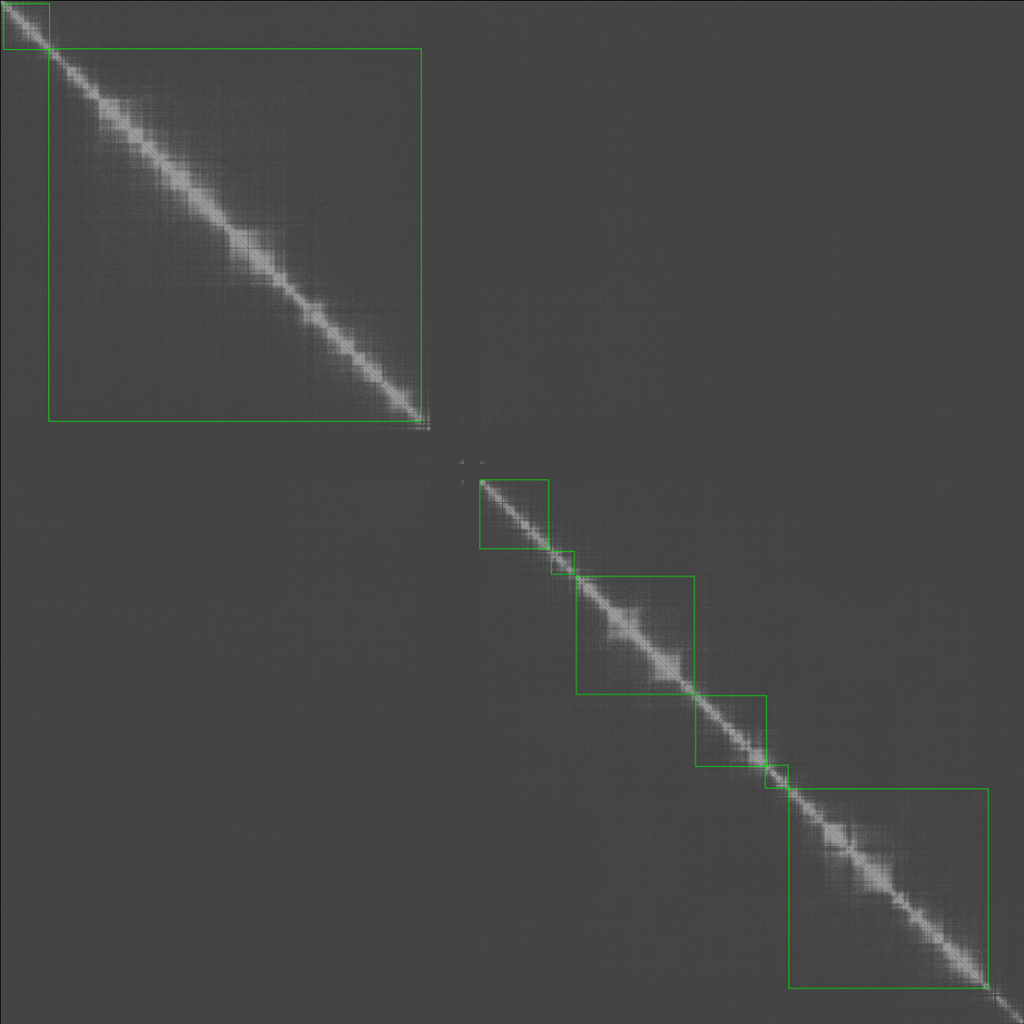


以上就是一個TAD calling的演算法發想,未來可以再加強它的效果,並與主流的TAD calling做比較
是一個對 “下一個世代” 的醫療科技充滿熱血的 Bioinformatican


